Abstract
Background: Erdheim-Chester disease (ECD) is a rare hematological malignancy, belonging to the L-group histiocytoses. ECD is characterized by multi-systemic proliferations of mature histiocytes in a background of inflammatory stroma. The inflammatory and neoplastic characteristics of the disease comprise a complex medical challenge for its diagnosis and treatment.
MicroRNAs (miRNAs/miRs) are short non-coding RNAs (~22 nucleotides) that regulate gene expression in a sequence specific manner and play an important role in cancer development and progression. Since miRNAs are released into the blood by tumor cells, they may be used as biomarkers to distinguish between cancer patients and healthy individuals and to assist in determining treatment response. Moreover, miRNA-mRNA interactions can determine the molecular mechanism by which miRNAs and their target genes are involved in ECD and may suggest novel therapeutic options for these patients. To date, this is the first study elucidating the role of miRNA in ECD.
Aims: The main focus of this study is to identify miRNAs that are differentially expressed in ECD patients compared to healthy controls and any clinical utility they have as potential biomarkers in ECD diagnosis, as well as to investigate their role in ECD pathogenesis, which may lead to new therapeutic options.
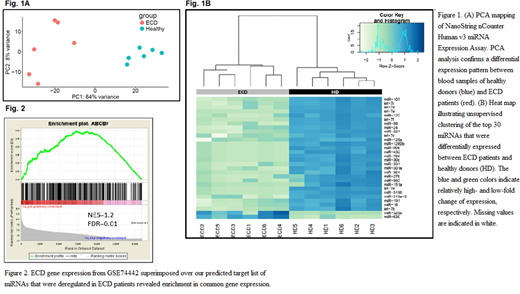
Preliminary results: Using the nCounter Human miRNA Expression Assay (NanoString Technologies), we analyzed the plasma miRNA expression profiles of 6 ECD patients (BRAF V600E) compared to 6 healthy individuals. Of the 800 mature miRNAs analyzed, 234 miRNAs showed different expression levels in these samples. Principal component analysis (PCA) was applied to experimental quality control. The miRNAs from healthy donors were clustered separately from the ECD samples indicating a distinct miRNA expression pattern between these groups (Fig. 1A, 1B). Among the 131 miRNAs remaining in the final analysis (FDR<0.05),110 miRNAs were downregulated in ECD patients compared to those of healthy controls, and 21 miRNAs were upregulated in ECD samples compared to those of the controls.
We validated the analysis method by quantitative real-time polymerase chain reaction (qRT-PCR) and found a positive correlation between miRs-15a, 16, 125a, 223, 21, 34a, 155 and miR-630 expression obtained by the NanoString array. This may indicate the potential use of miRNAs as biomarkers in ECD.
To determine potential target genes and signaling pathways implicated in ECD, we analyzed the predicted pathways of the top 30 downregulated miRNAs that were differentially expressed between the two groups using the Ingenuity® Pathway Analysis (IPA) and DIANA-miRPath v3.0 database. Reassuringly, the analysis identified cancer, inflammatory disease, and inflammatory response (p<0.01) as the main disease and disorder related with the miRNA expression pattern, as well as oncogenic pathways such as MAPK, PI3K-AKT, RAS, ErbB, Hippo, and mTOR as the main molecular pathways related to the differentially-expressed miRNAs (p<0.009). This finding suggests that low expression of miRNAs results in up regulation of target genes that participate in cell survival signaling. These augmented pathways may be inhibited by novel therapeutic treatments such as PI3K inhibitors, mTOR pathway inhibitors, and MEK inhibitors in ECD patients.
Next, we examined if there is any correlation between the predicted target genes of the miRNAs (obtained by IPA) and the experimentally validated gene expression pattern in ECD patients. To that end, we downloaded RNA-seq results of ECD patients from the GEO database (GSE74442 deposited by Diamond et al) and compared this list to our predicted miRNA targets in ECD patients, using Gene Set Enrichment Analysis (GSEA). We found a positive correlation between the gene expression reported in the literature and the predicted target of our deregulated miRNAs (Fig. 2), indicating that the predicted target genes are enriched in this data set, suggesting that the differentially expressed miRNAs might have a crucial role in the pathogenesis of ECD.
Conclusions: Our preliminary data highlight the unique inflammatory and neoplastic features characteristic of ECD. These deregulated miRNAs may highlight new candidate gene targets allowing for a better understanding of the molecular mechanisms underlying the development of ECD and propose novel therapeutic treatments for these patients.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal